Vergne-Vaxelaire C, Bordier F., Fossey A., Besnard-Gonnet M., Debard A., Mariage A., Pellouin V., Perret A., Petit J.-L., Stam M., Salanoubat M., Weissenbach J., de Berardinis V., Zaparucha A.
Adv. Syn. Catal. 2013, 355, 1763-1779
Abstract
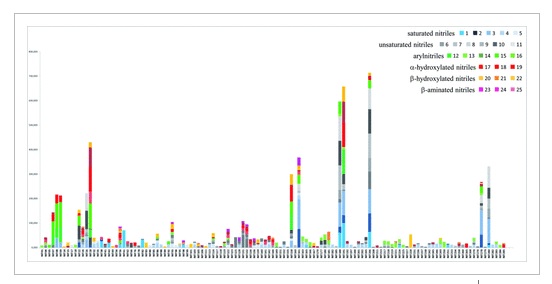
A high-throughput screening of candidate nitrilases against 25 structurally diverse substrates allowed us to create a wide collection of 125 experimentally validated nitrilases. The enzymes were selected by genomic approach from 700 diverse prokaryotic species and one metagenome as representative of the nitrilase family diversity. The enzymatic screening of this collection expands the biocatalytic toolbox for chemical synthesis by providing a large number of tested nitrilases with their assigned substrates. Three examples illustrate the synthetic potential of our enzyme collection. The syntheses of carboxylic acid building blocks, a β-substituted phenylpropanoic acid, a cyclic γ-keto carboxylic acid and a mononitrile monocarboxylic acid, were achieved from the corresponding nitrile substrates, using three new nitrilases (two from Sphingomonas wittichii and one from Syntrophobacter fumaroxidans). Improvements of nitrilase activities through the optimization of reaction parameters and the preparative biocatalytic synthesis are presented for these three examples.