Caparco A., A.Pelletier E., Petit JL., Jouenne A., Bommarius B., de Berardinis V., Zaparucha A., Champion J., Bommarius A., Vergne-Vaxelaire C.
Adv. Synth. Catal., 2020, 362(12), 2427-2436
Abstract
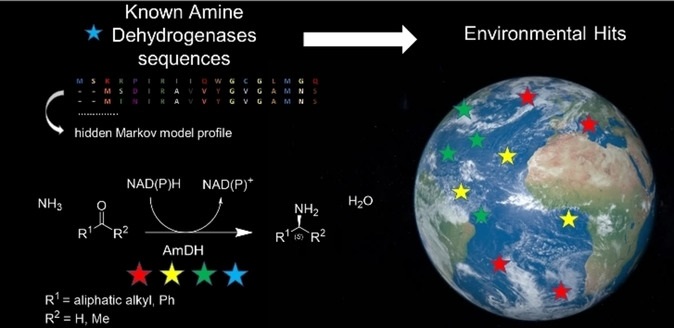
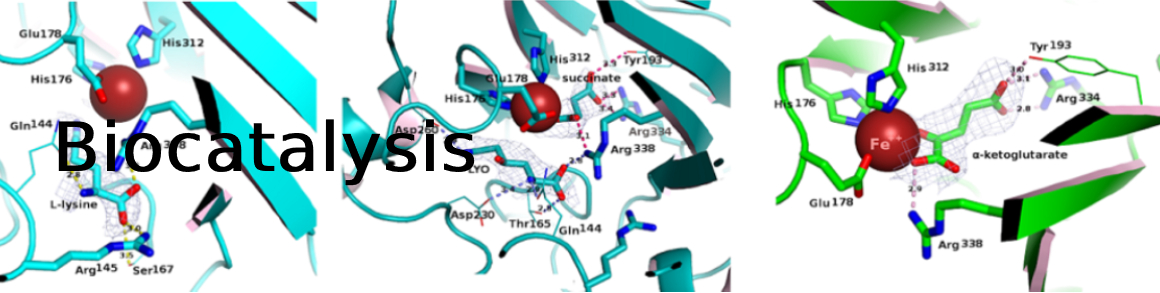
Amine dehydrogenases (AmDHs) catalyze the enzymatic reduction of ketones to amines, serving as a suitable biocatalytic route for amine synthesis. A limited number of experimentally validated native AmDHs (nat-AmDHs) have been reported recently, expanding the sequences with this function to complement the small set of engineered enzymes. Since researchers can now probe into the vast diversity of enzymes within niche environments by a metagenomics approach, a tandem metagenomic and bioinformatic approach is a powerful tool to identify new members of limited enzyme families to access new features in an iterative fashion. The previously untapped biocatalytic reservoirs of the ocean environment and human microbiome were screened for potential AmDHs using a hidden Markov model. Among the hundreds of hits, a subset of 18 enzymes was selected for further characterization and were confirmed to display AmDH activity. Additional analysis on six enzymes confirmed altered cofactor specificities and variation in substrate scopes, catalytic efficiencies, and active site residues compared to the reference nat-AmDHs previously described. Particularly, MATOUAmDH2 from an eukaryotic organism demonstrated specific activity of 11.07 and 0.88 U mg−1 toward isobutyraldehyde and 1,2-cyclohexadione respectively. Their abundance among the screened environments was also described. The protein sequence diversity of validated AmDHs reached by this metagenomics mining strategy highlights the success of such an approach. Metagenomically mined proteins, including eukaryotic ones, stand to increase the reach of biocatalysis towards enviromentally benign processes.